Theoretical Chemistry
The Theory Research Interest Group focuses on the development and application of theoretical and computational chemistry techniques. The group is highly interdisciplinary and collaborative, and is deeply engaged with the multi-department Lennard-Jones Centre for Computational Materials Science. Our aim is to advance fundamental understanding and computational tools in all areas of molecular science, and soft and condensed matter. Active research projects cover quantum dynamics and electronic structure, statistical mechanics, electrochemistry, machine learning, drug discovery, and energy landscapes from materials science to soft matter and biophysics.
Postgraduate applications
The Theory Group are always interested in receiving applications from highly qualified and talented candidates for postgraduate research. Find out more about our postgraduate study options and admissions process.
Modelling complex phenomena in nanoscale water flow (Image credit: Anna Bui, Cox group)
Snapshot of an amorphous carbon structure obtained from a machine learning potential of carving (Image credit: Patrick Rowe, Michaelides group)
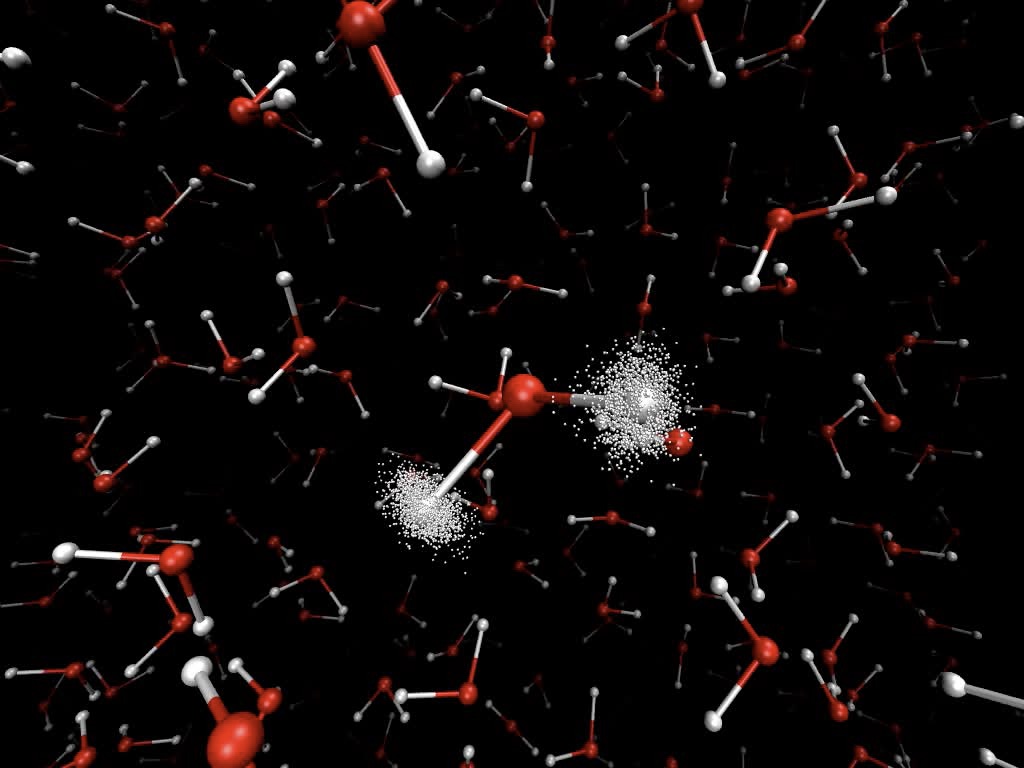

Quantum fluctuations of hydrogen atoms in ice, from a path-integral dynamics simulation carried out by George Trenins, Althorpe group.

